| Gene Symbol | erbb3 |
| Entrez Gene ID | 102290264 |
| Full Name | erb-b2 receptor tyrosine kinase 3 |
| Gene Type | protein-coding |
| Organism | Haplochromis burtoni(Burton's mouthbrooder) |
HOME
ORF » Species Summary » Haplochromis burtoni » erbb3 cDNA ORF clone
| Gene Symbol | erbb3 |
| Entrez Gene ID | 102290264 |
| Full Name | erb-b2 receptor tyrosine kinase 3 |
| Gene Type | protein-coding |
| Organism | Haplochromis burtoni(Burton's mouthbrooder) |
| mRNA | Protein | Name |
|---|---|---|
| XM_014340621.1 | XP_014196107.1 | receptor tyrosine-protein kinase erbB-3 |


Comparative transcriptomics of Eastern African cichlid fishes shows signs of positive selection and a large contribution of untranslated regions to genetic diversity.
Baldo L, Santos ME, Salzburger W
Genome biology and evolution3443-55(2011)
GeneRIFs: Gene References Into Functions What's a GeneRIF?
The following erbb3 gene cDNA ORF clone sequences were retrieved from the NCBI Reference Sequence Database (RefSeq). These sequences represent the protein coding region of the erbb3 cDNA ORF which is encoded by the open reading frame (ORF) sequence. ORF sequences can be delivered in our standard vector, pcDNA3.1+/C-(K)DYK or the vector of your choice as an expression/transfection-ready ORF clone. Not the clone you want? Click here to find your clone.
| CloneID | OHa28809 | |
| Clone ID Related Accession (Same CDS sequence) | XM_014340621.1 | |
| Accession Version | XM_014340621.1 Latest version! | Documents for ORF clone product in default vector |
| Sequence Information | ORF Nucleotide Sequence (Length: 4245bp) Protein sequence SNP |
|
| Vector | pcDNA3.1-C-(k)DYK or customized vector |  User Manual User Manual |
| Clone information | Clone Map |  MSDS MSDS |
| Tag on pcDNA3.1+/C-(K)DYK | C terminal DYKDDDDK tags | |
| ORF Insert Method | CloneEZ™ Seamless cloning technology | |
| Insert Structure | linear | |
| Update Date | 2015-09-28 | |
| Organism | Haplochromis burtoni(Burton's mouthbrooder) | |
| Product | receptor tyrosine-protein kinase erbB-3 | |
| Comment | Comment: MODEL REFSEQ: This record is predicted by automated computational analysis. This record is derived from a genomic sequence (NW_005179796.1) and transcript sequences (GBDH01028009.1 and JL523566.1) annotated using gene prediction method: Gnomon. Also see: Documentation of NCBI's Annotation Process ##Genome-Annotation-Data-START## Annotation Provider :: NCBI Annotation Status :: Full annotation Annotation Version :: Haplochromis burtoni Annotation Release 101 Annotation Pipeline :: NCBI eukaryotic genome annotation pipeline Annotation Software Version :: 6.4 Annotation Method :: Best-placed RefSeq; Gnomon Features Annotated :: Gene; mRNA; CDS; ncRNA ##Genome-Annotation-Data-END## ##RefSeq-Attributes-START## ab initio :: 1% of CDS bases assembly gap :: added 470 transcript bases to patch genome assembly gap ##RefSeq-Attributes-END## | |
1 | ATGAAGGTCC TTTTGAAGAA GGTGCAGCAG CAGGTGCTCT ACGTCCTGCT GCCGTGTGTA |
The stop codons will be deleted if pcDNA3.1+/C-(K)DYK vector is selected.
| RefSeq | XP_014196107.1 |
| CDS | 296..4540 |
| Translation |

Target ORF information:
Target ORF information:
|
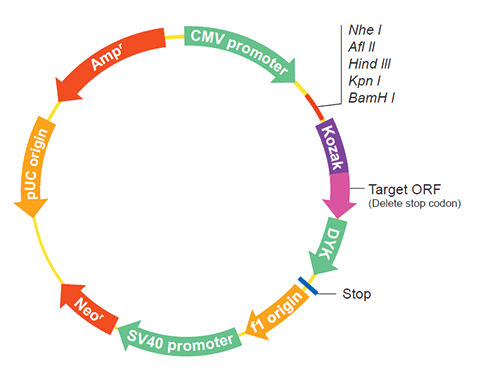 XM_014340621.1 XM_014340621.1 |

1 | ATGAAGGTCC TTTTGAAGAA GGTGCAGCAG CAGGTGCTCT ACGTCCTGCT GCCGTGTGTA |
The stop codons will be deleted if pcDNA3.1+/C-(K)DYK vector is selected.
Comparative transcriptomics of Eastern African cichlid fishes shows signs of positive selection and a large contribution of untranslated regions to genetic diversity. |