CRISPR 転写活性化SAMライブラリー
SAMライブラリーは、ゲノムワイドな機能獲得スクリーニングにおいて強力な転写活性化を誘導します。
SAMライブラリーは、ゲノム中の各コード領域に対して強力かつ特異的な転写活性化を誘導することで、ヒト細胞での機能獲得スクリーニングが可能です。CRISPR/Cas9
Synergistic Activation Mediator(SAM)は、内在性遺伝子の転写機能を強力に活性化するように設計されたタンパク質複合体です。CRISPR/Cas9 Synergistic
Activation
Mediator(SAM)は、内在性遺伝子の転写機能を強力に活性化するように設計されたタンパク質複合体です。SAMは、ガイドRNAによりゲノム中の特定の座位をターゲットするCas9ヌクレアーゼの特異性とリプログラミングのしやすさを利用しています。SAM複合体は3つの転写活性化ドメイン(VP64、P65、HSF1)を含んでおり、転写開始位置の200
bp上流内の部位をターゲットし転写を活性化します。SAMシステムは、切断活性を無くしたCas9を使用しており、内在性ゲノムのDNA鎖が切断されないことを確認しています。
ヒトのゲノムワイドSAMライブラリーは、RefSeqデータベース(23,430種のアイソフォーム)にある既知のコーディングアイソフォームすべてを活性化することができるため、遺伝子の活性化による機能獲得スクリーニングに有用です。このライブラリーは、ヒトおよびマウスの初代培養細胞、幹細胞、癌細胞、安定型発現細胞におけるスクリーニングに使用されています。SAMライブラリーを使用することにより、様々な条件下で細胞の生存に必須となる遺伝子や、癌細胞において薬剤耐性能を生じる遺伝子などを同定することが可能です。
SAMライブラリのデザインや構築に関する詳細は、以下の文献をご覧ください。Konermann S*, Brigham MD*, Trevino AE, Joung J,
Abudayyeh OO, Barcena C, Hsu PD, Habib N, Gootenberg JS, Nishimasu H, Nureki O & Zhang F. Genome-scale
transcriptional activation by an engineered CRISPR-Cas9 complex. Nature, doi:10.1038/nature14136。
SAMライブラリーの内容物
-
70,290種類のgRNA配列:
ヒトゲノム中の23,430種のユニークコーディングアイソフォームをターゲティングする各3種のsgRNA
genome。
-
Human SAM gRNA library is cloned into
pLenti
sgRNA(MS2)_zeoベクター骨格にクローニングされた、ヒトSAM gRNAライブラリー。
-
ライブラリーには、同時導入する必要がある、
pLenti
dCas-VP64_Blast および
pLenti
MS2-P-HSF1_Hygro プラスミドが添付されています。
SAMライブラリー納品物
すべてのSAMライブラリーは Industrial グレードとなります。発注量を増やすことも可能です。ライブラリーの追加オプションなどに関しては、お問い合わせください。
| Catalog # |
Library Name |
Vectors |
Amount |
Price |
|
SC1770-25
SC1770-100
SC1770-200
|
Human SAM Library |
|
25 µg
100 µg
200 µg
|
In Stock |
|
SC1754-25
SC1754-100
SC1754-200
|
|
25 µg
100 µg
200 µg
|
In Stock |
|
SC1844-25
SC1844-100
SC1844-200
|
Mouse SAM Library |
|
25 µg
100 µg
200 µg
|
In Stock |
機能獲得スクリーニングにおけるSAMライブラリーの使用法
SAMライブラリーは、ゲノムの転写開始点に隣接する上流域をターゲットとする各種CRISPR
gRNAをプールしたものです。SAMによる機能獲得遺伝子のスクリーニングには、ヒトのゲノムワイドSAM
sgRNAライブラリーの他に、2種類(dCas9-VP64およびMS2-P-HSF1)のSAM複合体を組み合わせて使用する必要があります。
さらに読む »
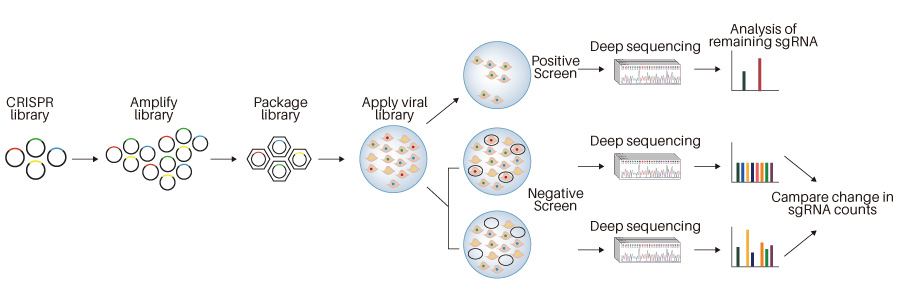
Cells must be transduced with all three SAM components. To enable co-transduction and
selection, all three lentiviral vectors have unique resistance markers. All three components are
delivered in lentiviral vectors optimized to produce high-titer virus for efficient lentiviral
transduction of primary cells or cultured cell lines. A cell population should be transduced with
the SAM library pool at a low MOI ensuring no more than 1 gRNA enters any given cell. After
transduction, deep sequencing with NGS should be performed to assess gRNA representation in the cell
pool before beginning a screening protocol. At the end of the screen, a second round of NGS is
performed, and data analysis should be performed to identify the guides that were lost or enriched
over the course of the screen. In order to identify true positive hits from a SAM library screen,
you should identify genes for which multiple guides were enriched. A more detailed SAM screening
protocol may be found on the
SAM / Genome
Engineering website.
SAMライブラリーの構成
SAM複合体は、3つのコンポーネントから構成されています:
-
〇 切断活性を無くしたCas9-VP64融合物
-
〇 テトラループとステムループに、二つのMS2 RNAアプタマーを組み込んだsgRNA
-
〇 MS2-P65-HSF1活性化ヘルパータンパク質
VP64、p65、HSF1の3つの異なる活性化ドメインを複合体に組み込むことで、相乗効果による強力な転写活性化を助けます。これらの3つのコンポーネントは、以下に示す通り、それぞれ別のレンチウイルスベクターにコード化されています。SAMが活性化されるには、これら3因子が同時に細胞に導入される必要があります。
ヒトのゲノムワイドなSAMガイドRNAライブラリーの配列については、csvファイルをご覧ください。